optuna.samplers.BaseSampler
- class optuna.samplers.BaseSampler[source]
Base class for samplers.
Optuna combines two types of sampling strategies, which are called relative sampling and independent sampling.
The relative sampling determines values of multiple parameters simultaneously so that sampling algorithms can use relationship between parameters (e.g., correlation). Target parameters of the relative sampling are described in a relative search space, which is determined by
infer_relative_search_space().The independent sampling determines a value of a single parameter without considering any relationship between parameters. Target parameters of the independent sampling are the parameters not described in the relative search space.
More specifically, parameters are sampled by the following procedure. At the beginning of a trial,
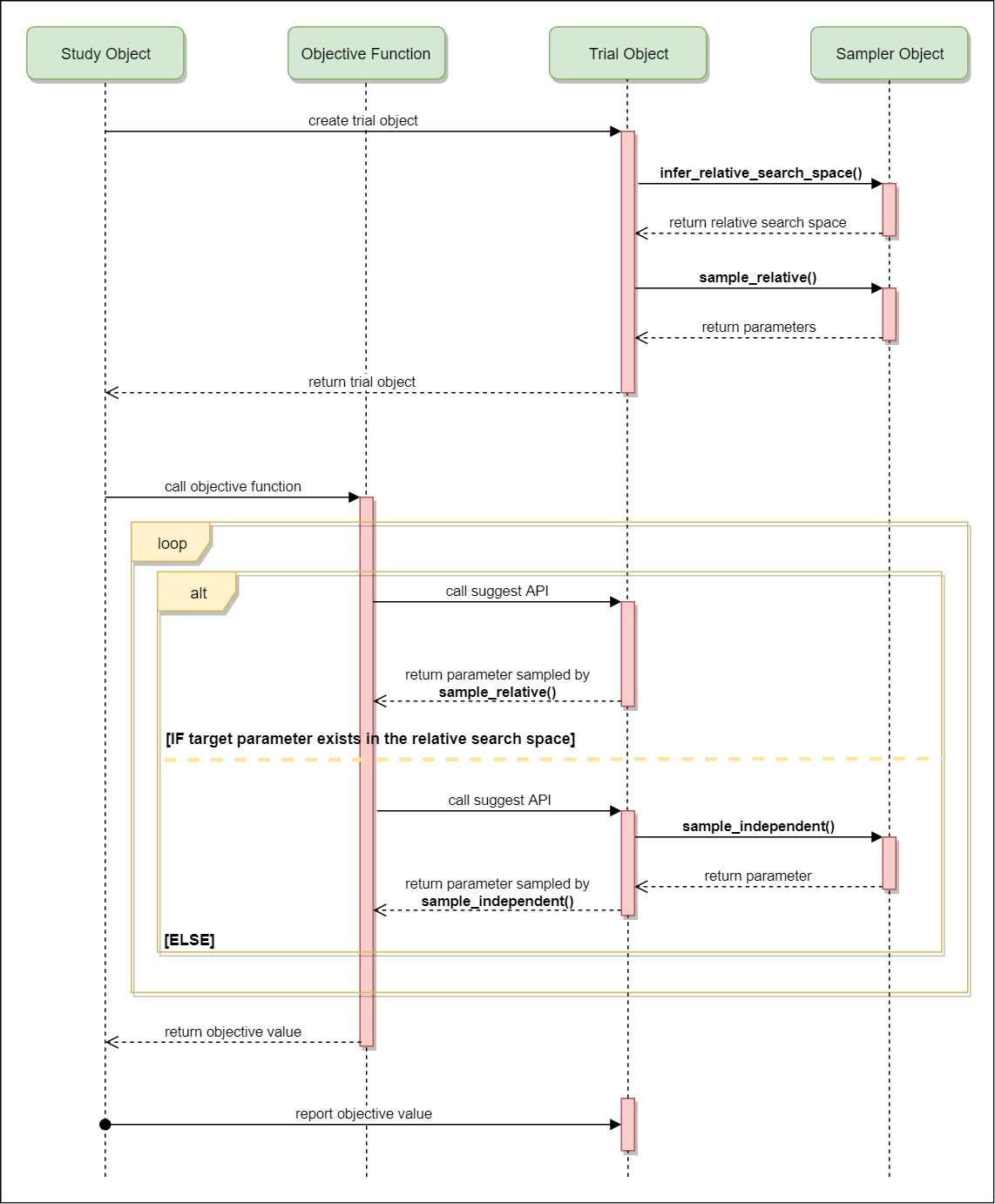
infer_relative_search_space()is called to determine the relative search space for the trial. During the execution of the objective function,sample_relative()is called only once when sampling the parameters belonging to the relative search space for the first time.sample_independent()is used to sample parameters that don’t belong to the relative search space.The following figure depicts the lifetime of a trial and how the above three methods are called in the trial.
Methods
after_trial(study, trial, state, values)Trial post-processing.
before_trial(study, trial)Trial pre-processing.
infer_relative_search_space(study, trial)Infer the search space that will be used by relative sampling in the target trial.
Reseed sampler's random number generator.
sample_independent(study, trial, param_name, ...)Sample a parameter for a given distribution.
sample_relative(study, trial, search_space)Sample parameters in a given search space.
- after_trial(study, trial, state, values)[source]
Trial post-processing.
This method is called after the objective function returns and right before the trial is finished and its state is stored.
Note
Added in v2.4.0 as an experimental feature. The interface may change in newer versions without prior notice. See https://github.com/optuna/optuna/releases/tag/v2.4.0.
- Parameters:
study (Study) – Target study object.
trial (FrozenTrial) – Target trial object. Take a copy before modifying this object.
state (TrialState) – Resulting trial state.
values (Optional[Sequence[float]]) – Resulting trial values. Guaranteed to not be
Noneif trial succeeded.
- Return type:
None
- before_trial(study, trial)[source]
Trial pre-processing.
This method is called before the objective function is called and right after the trial is instantiated. More precisely, this method is called during trial initialization, just before the
infer_relative_search_space()call. In other words, it is responsible for pre-processing that should be done before inferring the search space.Note
Added in v3.3.0 as an experimental feature. The interface may change in newer versions without prior notice. See https://github.com/optuna/optuna/releases/tag/v3.3.0.
- Parameters:
study (Study) – Target study object.
trial (FrozenTrial) – Target trial object.
- Return type:
None
- abstract infer_relative_search_space(study, trial)[source]
Infer the search space that will be used by relative sampling in the target trial.
This method is called right before
sample_relative()method, and the search space returned by this method is passed to it. The parameters not contained in the search space will be sampled by usingsample_independent()method.- Parameters:
study (Study) – Target study object.
trial (FrozenTrial) – Target trial object. Take a copy before modifying this object.
- Returns:
A dictionary containing the parameter names and parameter’s distributions.
- Return type:
Dict[str, BaseDistribution]
See also
Please refer to
intersection_search_space()as an implementation ofinfer_relative_search_space().
- reseed_rng()[source]
Reseed sampler’s random number generator.
This method is called by the
Studyinstance if trials are executed in parallel with the optionn_jobs>1. In that case, the sampler instance will be replicated including the state of the random number generator, and they may suggest the same values. To prevent this issue, this method assigns a different seed to each random number generator.- Return type:
None
- abstract sample_independent(study, trial, param_name, param_distribution)[source]
Sample a parameter for a given distribution.
This method is called only for the parameters not contained in the search space returned by
sample_relative()method. This method is suitable for sampling algorithms that do not use relationship between parameters such as random sampling and TPE.Note
The failed trials are ignored by any build-in samplers when they sample new parameters. Thus, failed trials are regarded as deleted in the samplers’ perspective.
- Parameters:
study (Study) – Target study object.
trial (FrozenTrial) – Target trial object. Take a copy before modifying this object.
param_name (str) – Name of the sampled parameter.
param_distribution (BaseDistribution) – Distribution object that specifies a prior and/or scale of the sampling algorithm.
- Returns:
A parameter value.
- Return type:
Any
- abstract sample_relative(study, trial, search_space)[source]
Sample parameters in a given search space.
This method is called once at the beginning of each trial, i.e., right before the evaluation of the objective function. This method is suitable for sampling algorithms that use relationship between parameters such as Gaussian Process and CMA-ES.
Note
The failed trials are ignored by any build-in samplers when they sample new parameters. Thus, failed trials are regarded as deleted in the samplers’ perspective.
- Parameters:
study (Study) – Target study object.
trial (FrozenTrial) – Target trial object. Take a copy before modifying this object.
search_space (Dict[str, BaseDistribution]) – The search space returned by
infer_relative_search_space().
- Returns:
A dictionary containing the parameter names and the values.
- Return type:
Dict[str, Any]