optuna.visualization.matplotlib.plot_parallel_coordinate
- optuna.visualization.matplotlib.plot_parallel_coordinate(study, params=None, *, target=None, target_name='Objective Value')[source]
Plot the high-dimensional parameter relationships in a study with Matplotlib.
Note that, if a parameter contains missing values, a trial with missing values is not plotted.
See also
Please refer to
optuna.visualization.plot_parallel_coordinate()for an example.Example
The following code snippet shows how to plot the high-dimensional parameter relationships.
import optuna def objective(trial): x = trial.suggest_float("x", -100, 100) y = trial.suggest_categorical("y", [-1, 0, 1]) return x ** 2 + y sampler = optuna.samplers.TPESampler(seed=10) study = optuna.create_study(sampler=sampler) study.optimize(objective, n_trials=10) optuna.visualization.matplotlib.plot_parallel_coordinate(study, params=["x", "y"])
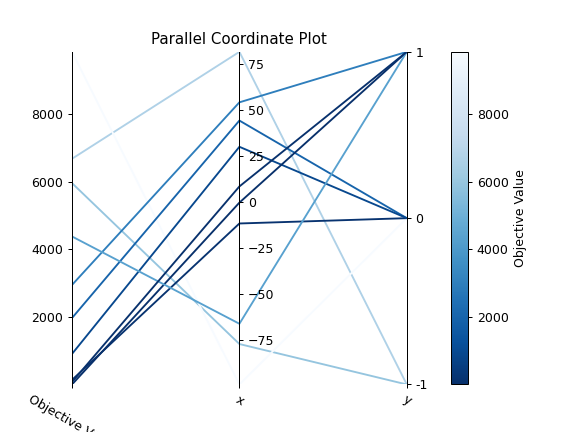
- Parameters:
study (Study) – A
Studyobject whose trials are plotted for their target values.params (list[str] | None) – Parameter list to visualize. The default is all parameters.
target (Callable[[FrozenTrial], float] | None) –
A function to specify the value to display. If it is
Noneandstudyis being used for single-objective optimization, the objective values are plotted.Note
Specify this argument if
studyis being used for multi-objective optimization.target_name (str) – Target’s name to display on the axis label and the legend.
- Returns:
A
matplotlib.axes.Axesobject.- Return type:
Note
The colormap is reversed when the
targetargument isn’tNoneordirectionofStudyisminimize.Note
Added in v2.2.0 as an experimental feature. The interface may change in newer versions without prior notice. See https://github.com/optuna/optuna/releases/tag/v2.2.0.